Talks
This page was produced as an assignment for Genetics 564 an undergraduate capstone course at UW-Madison.
Introduction
Cockayne Syndrome is a rare, autosomal recessive neurodegenerative disorder with a vast and complicated array of phenotypes. There are three types: Type 1, Type 2, and Type 3, and the phenotype of an affected individual varies both across and within type. All types are primarily characterized by an extreme photosensitivity. Some other common symptoms are cataracts, developmental delay (both mentally and physically), hearing problems, premature aging, microcephaly, and death.
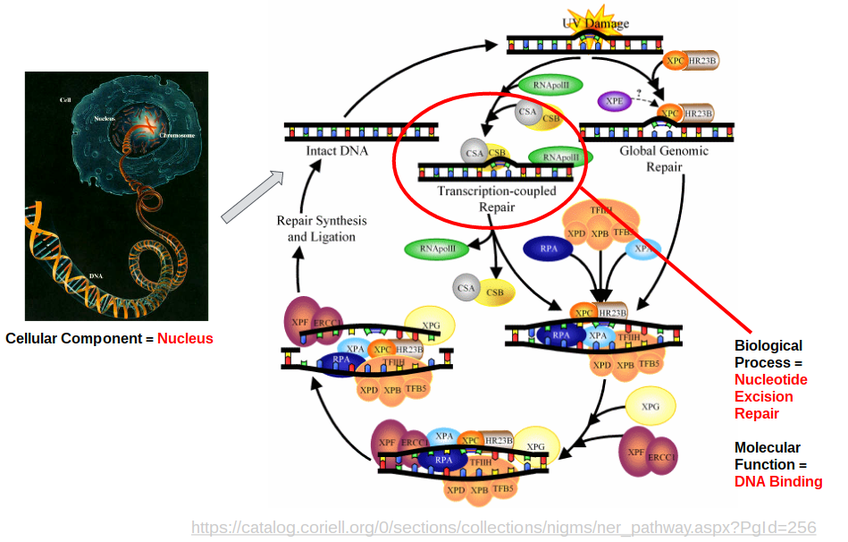
I will be focusing on Cockayne Syndrome Type 2, the congenital and most severe form of CS. The average age of death for a Type 2 patient is 7, compared to 12 for the less severe CS forms. CS Type 2 is caused by mutations in the ERCC6 gene (ERCC6 protein is sometimes called CSB). Figure 1 shows a detailed image of the pathway that ERCC6 is a crucial component of. ERCC6 is a protein that binds to damaged DNA in transcriptionally active genes and recruits other excision repair proteins to ultimately fix the DNA. When DNA is damaged, the RNA polymerase gets stuck and waits for ERCC6 to initiate the repair process. ERCC6's cellular component is the nucleus, it's biological process is nucleotide excision repair, and it's molecular function in DNA binding.
I will be focusing on Cockayne Syndrome Type 2, the congenital and most severe form of CS. The average age of death for a Type 2 patient is 7, compared to 12 for the less severe CS forms. CS Type 2 is caused by mutations in the ERCC6 gene (ERCC6 protein is sometimes called CSB). Figure 1 shows a detailed image of the pathway that ERCC6 is a crucial component of. ERCC6 is a protein that binds to damaged DNA in transcriptionally active genes and recruits other excision repair proteins to ultimately fix the DNA. When DNA is damaged, the RNA polymerase gets stuck and waits for ERCC6 to initiate the repair process. ERCC6's cellular component is the nucleus, it's biological process is nucleotide excision repair, and it's molecular function in DNA binding.
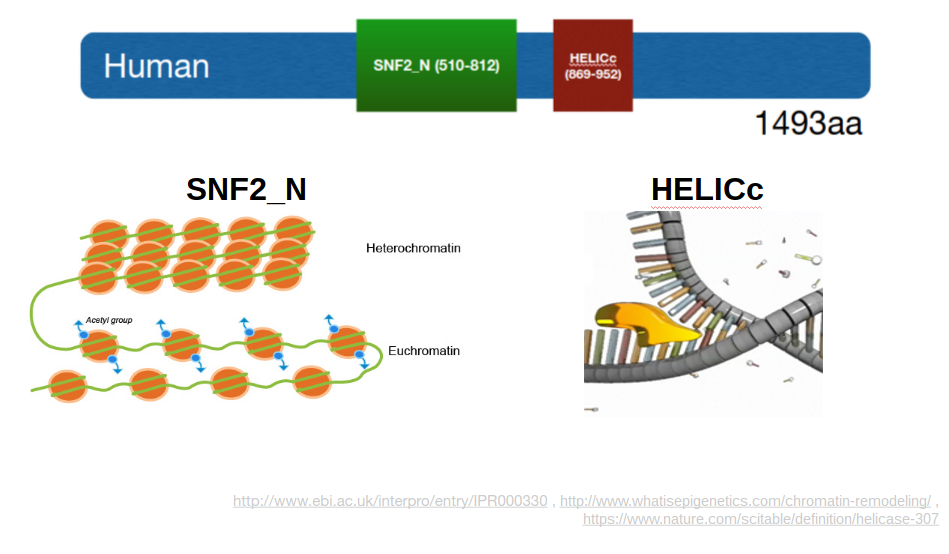
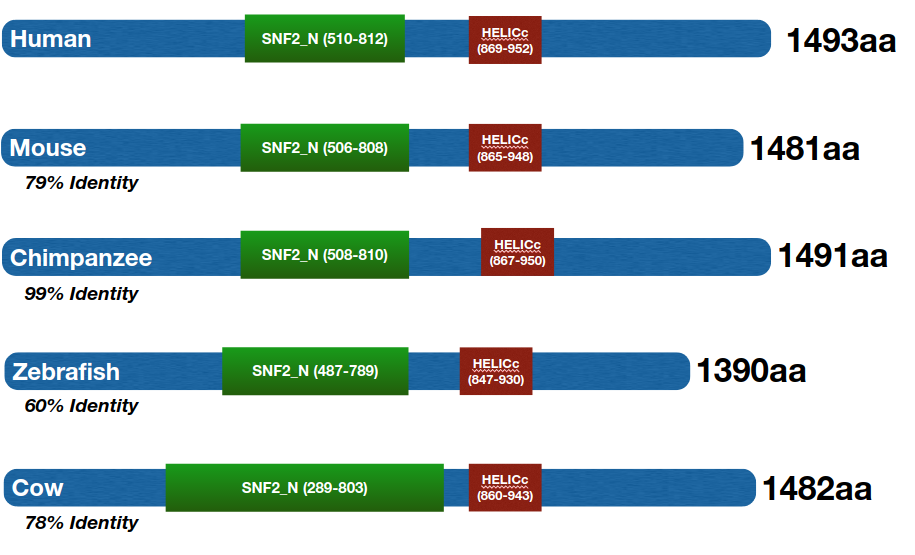
ERCC6 has two major protein domains: the HELICc domain and the SNF2_N domain (Figure 2). These domains are well conserved across ERCC6 homologs (Figure 3). The HELICc domain has helicase activity, and the SNF2_N has chromatin remodeling activity.
One of the primary phenotypes found in CS Type 2 patients is premature aging. Aging is thought to be caused by a build up of oxidative damage. Photosensitivity is a hallmark of CS because UV light causes DNA lesions in DNA. Oxidative stress and reactive oxygen species in the body also cause DNA damage, and I believe that ERCC6 likely interacts with oxidative damage in some way similar to its interaction with DNA lesions from UV light. However, the role in ERCC6 in regulating oxidative damage is not clear and has not been thoroughly investigated.
I hypothesize that ERCC6 has a DNA binding and lesion recognition role in the oxidative damage repair pathway, and ERCC6 loss leads to the aging phenotype in Cockayne Syndrome Type 2 patients because oxidative damage cannot be regulated. My primary goal in investigating this hypothesis is to determine how ERCC6 regulates DNA damage repair after oxidative stress associated with aging, and my overall long term goal is to better understand the role of ERCC6 in aging.
To explore this hypothesis, I have devised 3 Specific Aims that I feel will help address ERCC6's role in oxidative damage repair. I will carry out all experiments on mice, because they show a similar progeria-like premature aging phenotype to humans, in addition to having similar protein interaction networks to humans as well as conserved ERCC6 protein domains.
I hypothesize that ERCC6 has a DNA binding and lesion recognition role in the oxidative damage repair pathway, and ERCC6 loss leads to the aging phenotype in Cockayne Syndrome Type 2 patients because oxidative damage cannot be regulated. My primary goal in investigating this hypothesis is to determine how ERCC6 regulates DNA damage repair after oxidative stress associated with aging, and my overall long term goal is to better understand the role of ERCC6 in aging.
To explore this hypothesis, I have devised 3 Specific Aims that I feel will help address ERCC6's role in oxidative damage repair. I will carry out all experiments on mice, because they show a similar progeria-like premature aging phenotype to humans, in addition to having similar protein interaction networks to humans as well as conserved ERCC6 protein domains.
Specific Aims
Aim #1
|
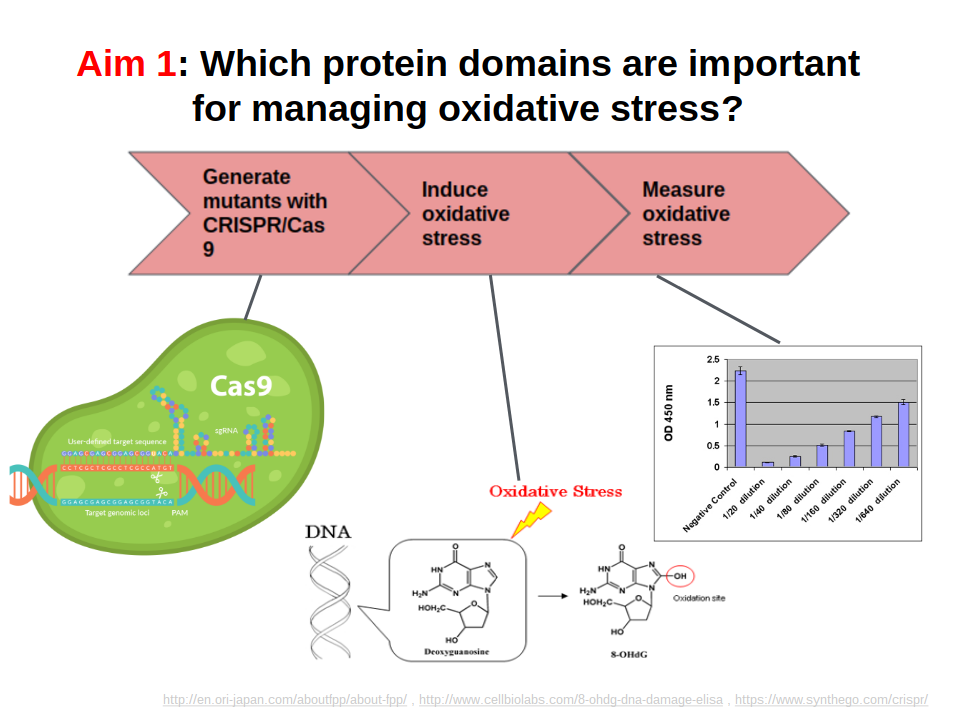
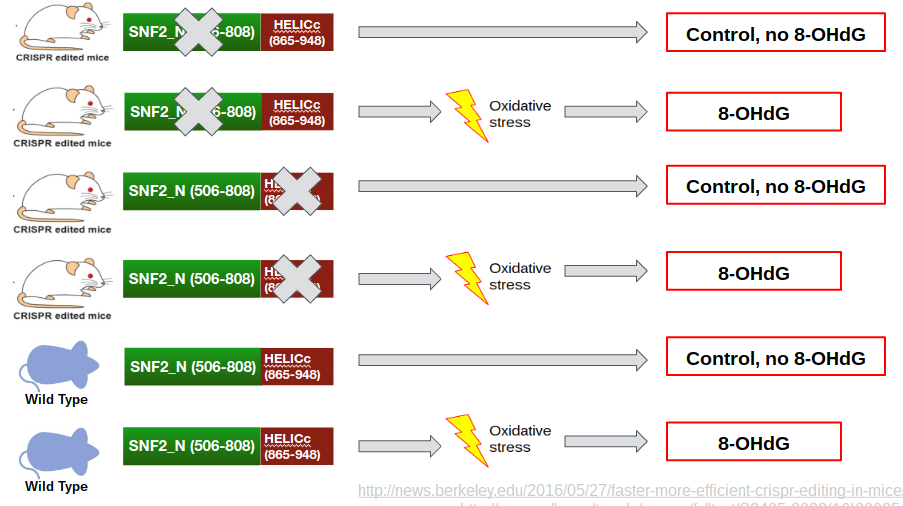
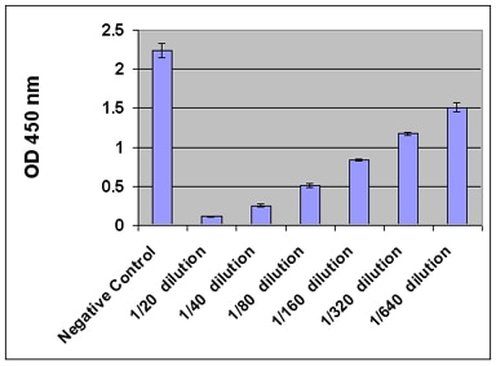
The purpose of Aim #1 is to identify which of the 2 ERCC6 protein domains is more important in mitigating oxidative damage. This is done using knockout mice for each domain, as well as a control. The experimental design for Aim #1 is shown in Aim Figure #1. There are two domains of ERCC6, the SNF2_N domain and the HELICc domain. HELICc has helicase activity, while SNF2_N has chromatin remodeling activity. These domains were identified using SMART and Pfam, and the domains are knocked out using CRISPR/Cas9. An oxidative derivative of deoxyguanosine, called 8-OHdG, is measured using the Cell Biolabs 8-OHdG ELISA after oxidative stress has been induced. 8-OHdG is processed by the body and excreted in urine in healthy individuals. I hypothesize that because ERCC6 domain knockout mice do not have a fully functioning ERCC6 protein, levels of oxidative damage will be higher in these mice and 8-OHdG will not properly be processed. Therefore, I hypothesize that levels of excreted 8-OHdG in urine will be lower in domain knockouts relative to the controls, and that it will specifically be lowest in the HELICc domain, because this domain is the integral domain that allows ERCC6 to bind to damaged DNA. An example output of the Cell Biolabs 8-OHdG ELISA can be seen in Aim Figure #2.
|
Aim #2
|
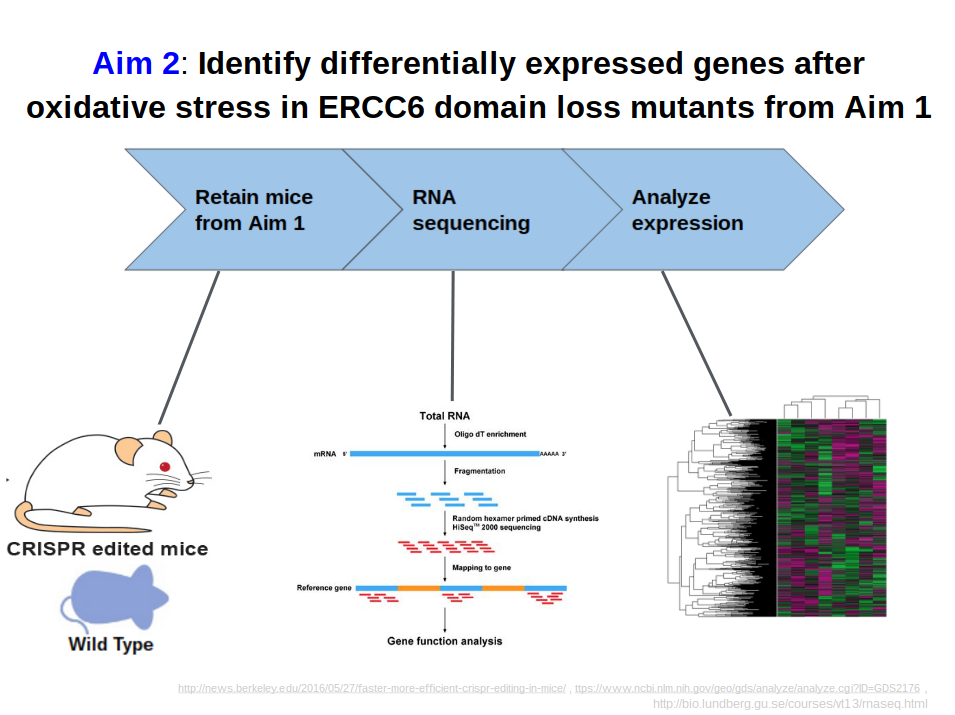
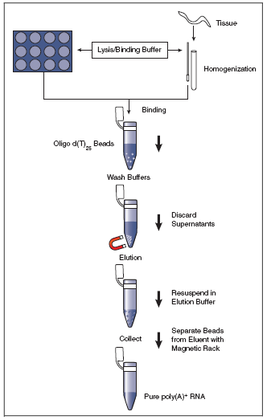
The purpose of Aim #2 is to identify overall global gene expression changes between the WT and domain knockout mice from Aim #1 after oxidative damage is induced. Just as in Aim #1, I will have a WT mouse line and 2 domain knockout mouse lines, and there will be 2 mice for each of these categories, one with and one without oxidative damage. I will isolate RNA with the New England Biolabs Magnetic mRNA Isolation Kit and will do RNA-sequencing to generate gene expression profiles for both mice in all 3 categories. The RNA isolation procedure is shown in Aim Figure #3, and is summarized on the webpage for the product. The profiles will be sorted and visualized by GO terms with PANTHER, and processes that are affected by the different domain knockouts will be identified. I hypothesize that the genes that are downregulated in the HELICc domain knockout will be genes in biological processes such as "aging" or "oxidative stress response".
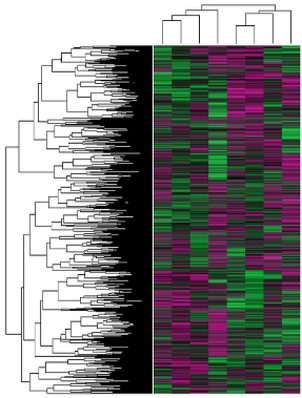
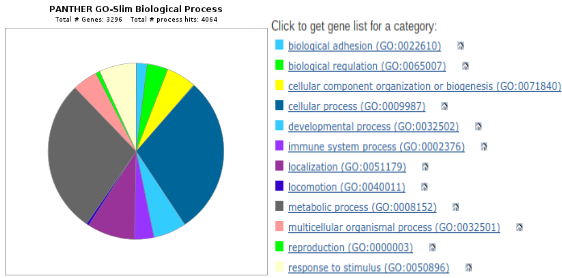
An example gene expression profile, also known as a heat map, can be seen in Aim Figure #4. The analysis of this heat map using PANTHER and sorting by GO terms can be seen in Aim Figure #5. |
Aim #3
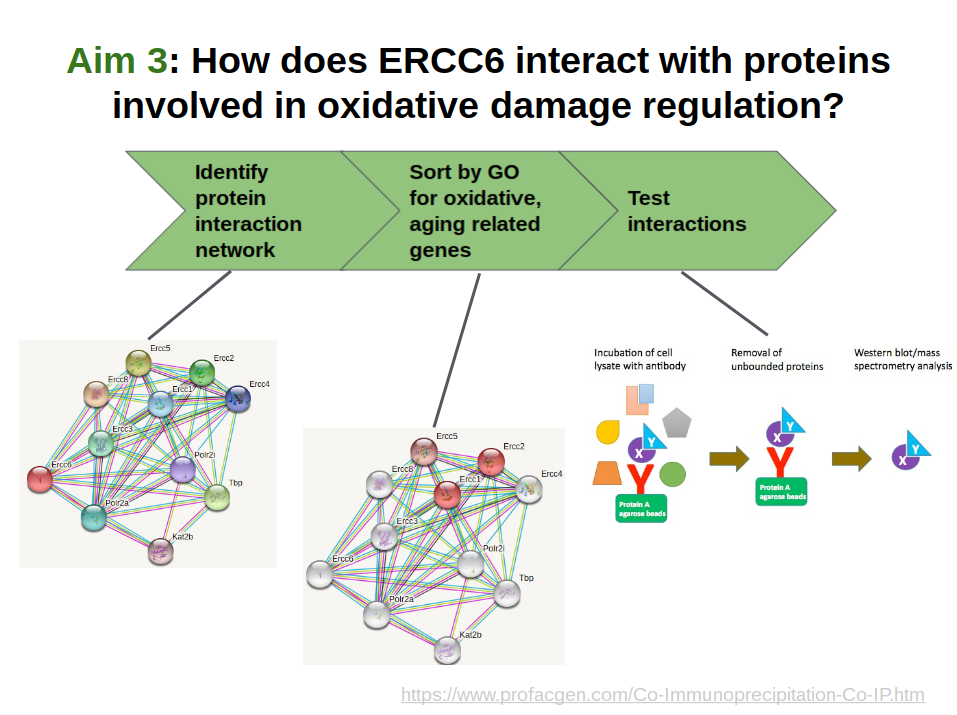
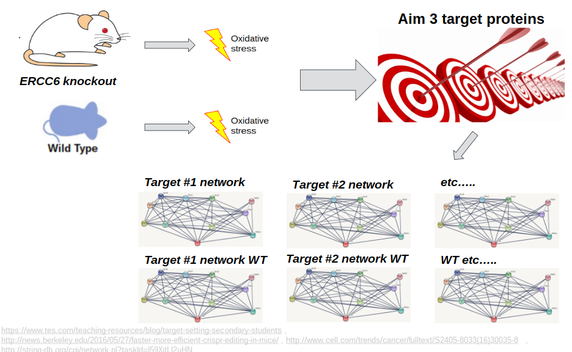
The purpose of Aim #3 is to confirm genes that directly bind with ERCC6 in mice and then observe ERCC6's interaction with these proteins in mice under oxidative stress. I hypothesize that the target proteins that directly bind with ERCC6 in the presence of oxidative damage will be significantly affected by it's loss, and that these proteins are likely downregulated in Cockayne Syndrome patients. I also believe that these proteins will be crucial players in the oxidative repair pathway, so ERCC6 loss significantly decreases the efficacy of this pathway in CS patients because of the lost interaction with ERCC6, ultimately resulting in the premature aging phenotype. The goal of this aim is to identify these proteins with the hope of finding potential targets for therapeutic medicine.
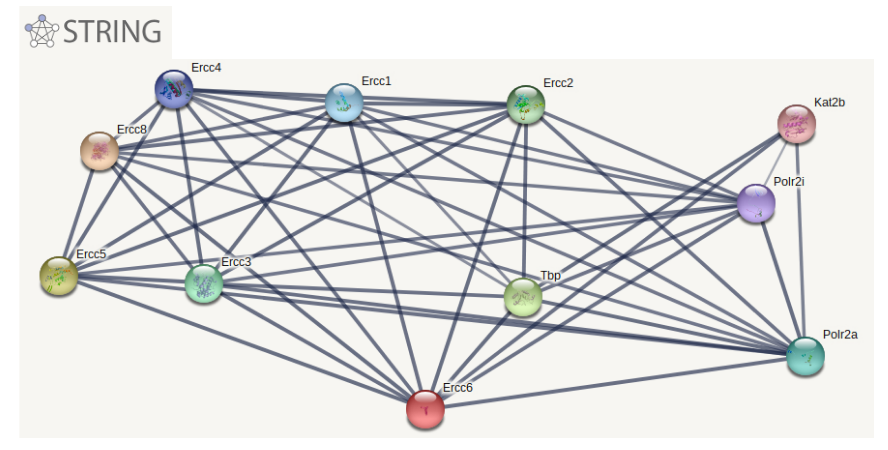
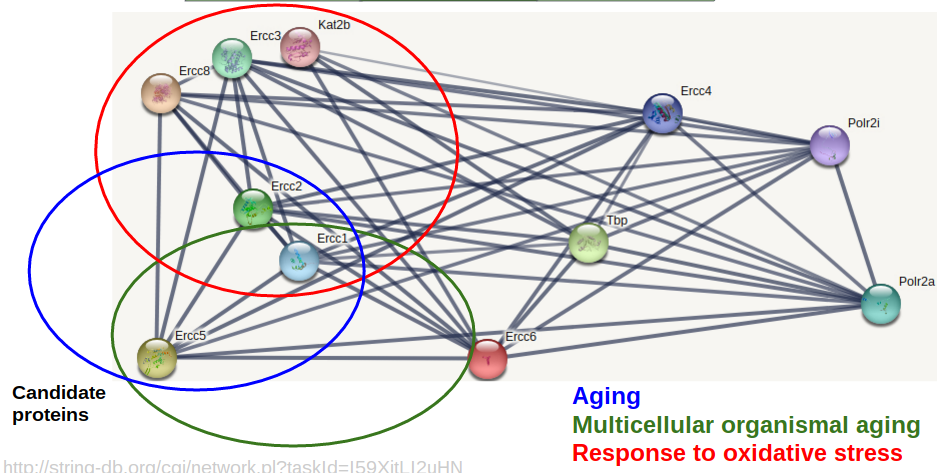
First, I will identify ERCC6's protein interaction network in mice using the STRING database. The network can be seen in Aim Figure #6. Then I will sort the interaction network by GO terms related to aging and oxidative stress response. A crude sorting can be seen in Aim Figure #7. The genes which have these GO terms will be my candidate genes. I will induce oxidative stress in a WT mouse and do co-immunoprecipitation to test ERCC6 binding to the candidate genes in the oxidative stress mouse and in a WT mouse without oxidative stress. I will use Western Blotting to analyze the protein binding. The proteins that bind to ERCC6 will be the target genes for further analysis as targets for medicine for CS patients.
First, I will identify ERCC6's protein interaction network in mice using the STRING database. The network can be seen in Aim Figure #6. Then I will sort the interaction network by GO terms related to aging and oxidative stress response. A crude sorting can be seen in Aim Figure #7. The genes which have these GO terms will be my candidate genes. I will induce oxidative stress in a WT mouse and do co-immunoprecipitation to test ERCC6 binding to the candidate genes in the oxidative stress mouse and in a WT mouse without oxidative stress. I will use Western Blotting to analyze the protein binding. The proteins that bind to ERCC6 will be the target genes for further analysis as targets for medicine for CS patients.
Future Directions
|
Moving forward, I propose creating a full ERCC6 knockout mouse, and inducing oxidative stress in this mouse and in a WT. I will analyze the protein interaction networks for each target protein identified in Aim #3 for both the mice, and look for differences between WT and ERCC6 knockout in the interactions between the target proteins and their respective interaction networks. A visualization of this experiment can be seen in Figure X. This will provide a more global understanding of the effects of ERCC6 loss on biological processes other than aging within the body, with the goal of hopefully understanding the complexity of the vast CS phenotypes in more detail.
|
Final talk download
| beethemfinal_42717.pptx.pdf | |
| File Size: | 3356 kb |
| File Type: | |
Past talk drafts
| beethemdraft2_4417.pptx.pdf | |
| File Size: | 1016 kb |
| File Type: | |
| beethemdraft2_4417.pptx.pptx | |
| File Size: | 1613 kb |
| File Type: | pptx |
| beethemdraft1_22117.pdf | |
| File Size: | 781 kb |
| File Type: | |
This website was created for Genetics 564 by Zachary Beethem, an undergraduate genetics major at UW-Madison.
He can be reached via email: [email protected]
Date of last website update: May 2017
He can be reached via email: [email protected]
Date of last website update: May 2017