Gene Ontology
This page was produced as an assignment for Genetics 564 an undergraduate capstone course at UW-Madison.
What is gene ontology?
Gene ontology is a way of standardizing genetic terminology with the purpose of creating a consistent vocabulary for use in describing gene products across studies and databases [1]. The volume of published genetic and molecular biology research and the huge amount of ongoing research calls for a way of standardizing and consolidating the jargon used to relay pertinent information. The Gene Ontology Consortium's GO database is one database that attempts to standardize biological information, and it was used to explore the gene ontology the ERCC6 gene product, CSB. This database uses 3 categories to organize information. These categories are molecular function, cellular component, and biological process.
Molecular Function
|
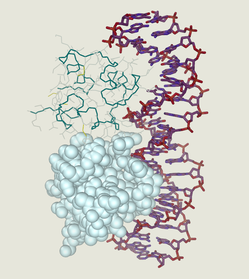
Molecular function describes the biochemical reactions carried out by the gene product [2]. Molecular function is where the activity of the product is detailed. For example, if a gene product were some sort of ligand, then one of the GO terms used in the molecular function category would be 'binding'.
ERCC6's molecular functions are DNA binding, protein binding, and chromatin binding [3,4]. |
Cellular Component
|
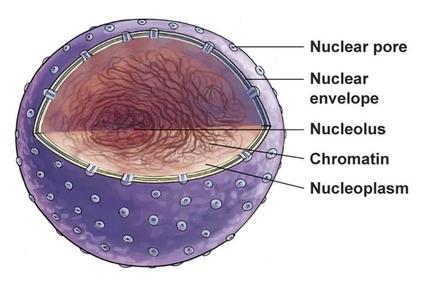
Cellular component describes where in a cell the molecular function is taking place. Often a gene product's cellular component is determined because the product has been found there, so the molecular function is assumed to also take place in the same area of the cell [2].
ERCC6's cellular components are the nucleoplasm, nucleus, nucleolus, and transcription elongation factor complex [3,4]. This makes sense because ERCC6 works in nucleotide excision repair during transcription, which takes place inside the nucleus. |
Biological Process
|
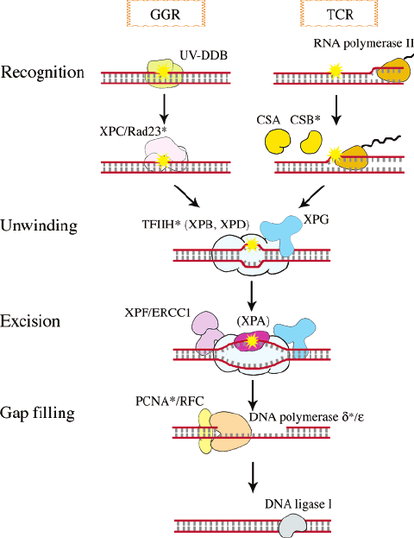
Biological process describes the product's overall objective within a cell [2]. Processes are the pathways where a product's molecular functions are utilized to achieve some goal in the cell. These processes are where the gene product goes to work.
ERCC6's biological processes include [3,4]:
ERCC6 has many biological processes that it is associated with. For Cockayne Syndrome, it's roles in response pathways and nucleotide excision repair are altered, leading to the complicated CS phenotype. |
Conclusions
|
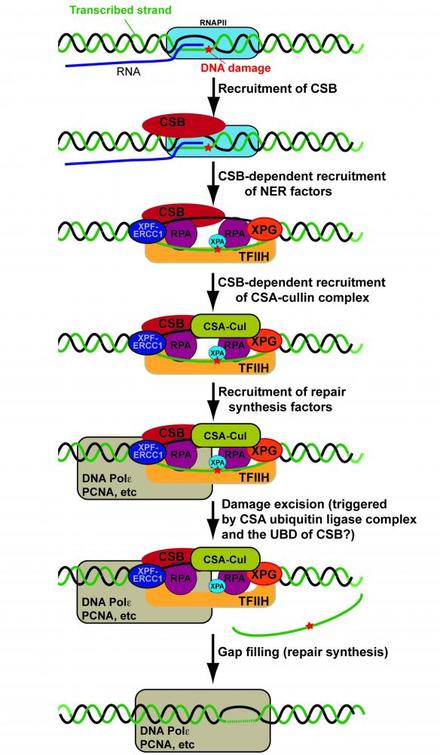
ERCC6 has ontologies that make sense given the known roles of the gene product CSB. ERCC6 plays a vital role in transcription-coupled nucleotide excision repair, which takes place simultaneously with transcription inside the nucleus. It helps fix DNA lesions during transcription, and is critical in recognition of these lesions. As a result, it needs to be able to bind DNA, which is one of it's molecular functions. In addition, CSB recruits other excision repair proteins, which requires the ability to bind proteins, as shown in Figure 4. ERCC6 also has some ability to alter chromatin structure in an ATP-dependent manner, which explains its chromatin molecular function ontology [5].
ERCC6's biological processes ontologies also make sense, given the known roles of CSB in the cell. Transcription coupled nucleotide excision repair fixes DNA lesions that result from factors such as UV light and radiation, and ERCC6's biological processes include responses to these factors. ERCC6's role in DNA repair processes are clear, and it's molecular function and cellular component ontologies corroborate it's involvement in the identified biological processes. |
References
[1] Doms, A. & Schroeder, M. (2005). GoPubMed: exploring PubMed with the Gene Ontology. Nucleic Acids Res, 33(2): W783-W786. doi: 10.1093/nar/gki470 https://academic.oup.com/nar/article/33/suppl_2/W783/2505675/GoPubMed-exploring-PubMed-with-the-Gene-Ontology
[2] Hill, D. P., Berardini, T. Z., Howe, D. G., & Auken, K. M. (2009). Representing ontogeny through ontology: A developmental biologist's guide to the gene ontology. Molecular Reproduction and Development. doi:10.1002/mrd.21130 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2830379/
[3] Gene Ontology Consortium. http://geneontology.org/
[4] Uniprot. http://www.uniprot.org/uniprot/Q03468
[5] Lake, R. J., Boetefuer, E. L., Won, K.-J., & Fan, H.-Y. (2016). The CSB chromatin remodeler and CTCF architectural protein cooperate in response to oxidative stress. Nucleic Acids Research, 44(5), 2125–2135. http://doi.org/10.1093/nar/gkv1219 #0
Images and Videos
Cover image: http://www.huffingtonpost.com/2013/04/14/human-gene-supreme-court_n_3081399.html
Figure 1: https://en.wikipedia.org/wiki/DNA-binding_protein
Figure 2: https://www.google.com/search?
q=nucleoplasm&rlz=1CAACAO_enUS698US699&espv=2&source=lnms&tbm=isch&sa=X&ved=0ahUKEwjux77u89LSAhXJslQKHS25Ce8Q_AUIBigB&biw=1920&bih=942#imgrc=bY6zyHqklcIwIM:
Figure 3: https://www.researchgate.net/figure/7310454_fig3_Figure-4-Nucleotide-excision-repair-NER-NER-pathways-can-be-classified-into-two
Figure 4: https://www.crick.ac.uk/research/a-z-researchers/researchers-p-s/jesper-svejstrup/projects/
This website was created for Genetics 564 by Zachary Beethem, an undergraduate genetics major at UW-Madison.
He can be reached via email: [email protected]
Date of last website update: April 2017
[1] Doms, A. & Schroeder, M. (2005). GoPubMed: exploring PubMed with the Gene Ontology. Nucleic Acids Res, 33(2): W783-W786. doi: 10.1093/nar/gki470 https://academic.oup.com/nar/article/33/suppl_2/W783/2505675/GoPubMed-exploring-PubMed-with-the-Gene-Ontology
[2] Hill, D. P., Berardini, T. Z., Howe, D. G., & Auken, K. M. (2009). Representing ontogeny through ontology: A developmental biologist's guide to the gene ontology. Molecular Reproduction and Development. doi:10.1002/mrd.21130 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2830379/
[3] Gene Ontology Consortium. http://geneontology.org/
[4] Uniprot. http://www.uniprot.org/uniprot/Q03468
[5] Lake, R. J., Boetefuer, E. L., Won, K.-J., & Fan, H.-Y. (2016). The CSB chromatin remodeler and CTCF architectural protein cooperate in response to oxidative stress. Nucleic Acids Research, 44(5), 2125–2135. http://doi.org/10.1093/nar/gkv1219 #0
Images and Videos
Cover image: http://www.huffingtonpost.com/2013/04/14/human-gene-supreme-court_n_3081399.html
Figure 1: https://en.wikipedia.org/wiki/DNA-binding_protein
Figure 2: https://www.google.com/search?
q=nucleoplasm&rlz=1CAACAO_enUS698US699&espv=2&source=lnms&tbm=isch&sa=X&ved=0ahUKEwjux77u89LSAhXJslQKHS25Ce8Q_AUIBigB&biw=1920&bih=942#imgrc=bY6zyHqklcIwIM:
Figure 3: https://www.researchgate.net/figure/7310454_fig3_Figure-4-Nucleotide-excision-repair-NER-NER-pathways-can-be-classified-into-two
Figure 4: https://www.crick.ac.uk/research/a-z-researchers/researchers-p-s/jesper-svejstrup/projects/
This website was created for Genetics 564 by Zachary Beethem, an undergraduate genetics major at UW-Madison.
He can be reached via email: [email protected]
Date of last website update: April 2017