Interaction Networks
This page was produced as an assignment for Genetics 564 an undergraduate capstone course at UW-Madison.
What are interaction networks?
Interaction networks are a way to visualize large scale protein-protein interactions. Protein interaction networks offer a way to understand and visualize a protein's relationship to other proteins and create a biological network to help deduce a protein's role in a cell [1]. Deduction of protein function also extends to previously unknown or characterized proteins, because one can discover the general biological processes that the unknown protein is involved in [2]. To study the interaction networks of ERCC6, the online database STRING was used.
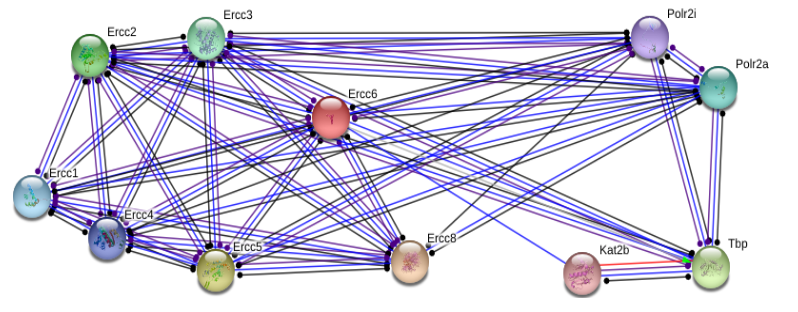
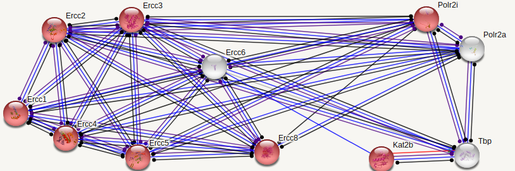
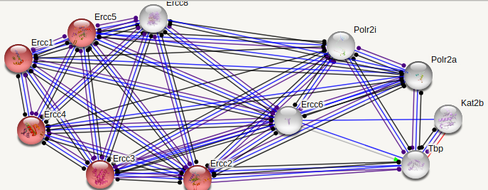
ERCC6 interaction networks in Mus musculus
|
|
|
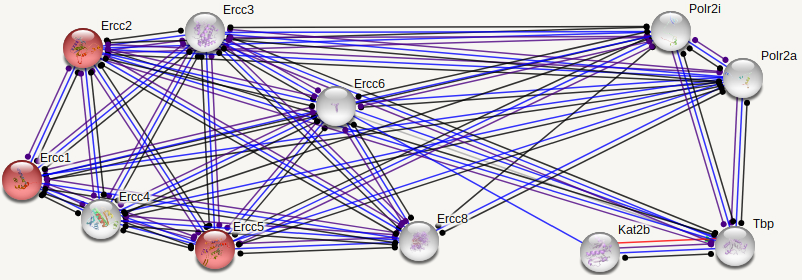
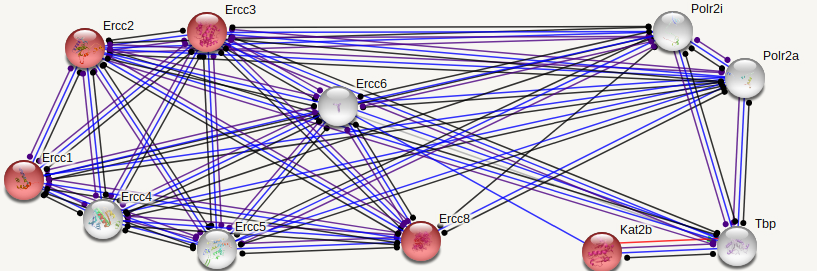
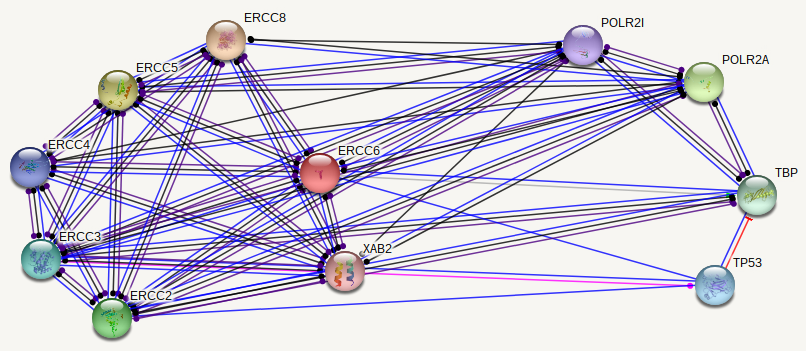
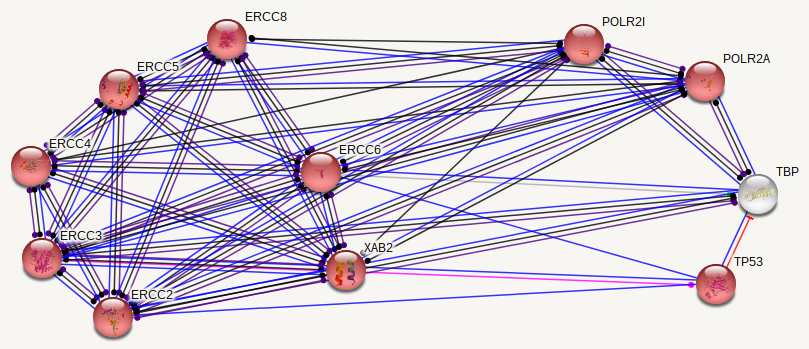
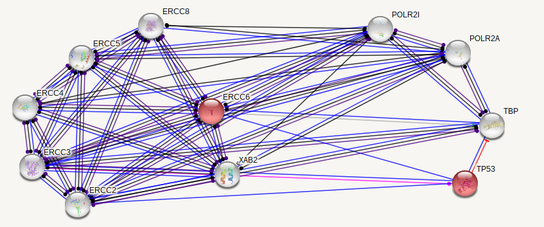
ERCC6 interaction networks in Homo sapiens
Analysis
The interaction networks between humans and mice show similarities, but also a few significant differences. Both interaction networks had the same two parameters (confidence above 0.900 and show 10 of the first shell), yet some different proteins and differences in GO terms appeared. The human ERCC6 network had two proteins that the mouse network did not: XAB2, a transcription-coupled repair binding protein, and TP53, tumor protein p53. The mouse network had Kat2b, a histone acetyltransferase, and ERCC1, an endonuclease responsible for the 5' incision during DNA repair. In humans, all but one protein (TBP) are involved in nucleotide excision repair, while 5 of the proteins in mice are involved in nucleotide excision repair. Perhaps most interesting for the analysis of ERCC6's role in oxidative damage repair are the proteins found in the mice network that have biological process GO terms of "aging" and "response to oxidative stress". The presence of these GO terms suggests a possible association between ERCC6 and oxidative damage repair, likely through these proteins found in the interaction network.
References
[1] Vazquez, A., Flammini, A., Maritan, A., Vespignani, A. (2003). Global protein function prediction from protein-protein interaction networks. Nature Biotechnology, 21: 697-700. doi:10.1038/nbt825
[2] Goehler, H., Lalowski, M., Stelzl, U., et al. (2005). A protein interaction network links GIT1, an enhancer of huntingtin aggregation, to Huntington's disease. Molecular Cell, 19(2): 287. http://doi.org/10.1016/j.molcel.2004.09.016
[3] STRING. http://string-db.org/cgi/input.pl?UserId=pr2yjwXlURmS&sessionId=rLd6tI7IrE5T&input_page_show_search=on
Images and Videos
Cover image: https://www.mdc-berlin.de/16074687/en/news/archive/2008/20080910-erwin_schr_dinger_prize_2008_goes_to_resea/P__PRESS2008_WANKER_SCHROEDINGER_PRIZE_PROTEIN_NETWORK.jpg
Interaction network images: All come from a STRING session.
This website was created for Genetics 564 by Zachary Beethem, an undergraduate genetics major at UW-Madison.
He can be reached via email: [email protected]
Date of last website update: April 2017
[1] Vazquez, A., Flammini, A., Maritan, A., Vespignani, A. (2003). Global protein function prediction from protein-protein interaction networks. Nature Biotechnology, 21: 697-700. doi:10.1038/nbt825
[2] Goehler, H., Lalowski, M., Stelzl, U., et al. (2005). A protein interaction network links GIT1, an enhancer of huntingtin aggregation, to Huntington's disease. Molecular Cell, 19(2): 287. http://doi.org/10.1016/j.molcel.2004.09.016
[3] STRING. http://string-db.org/cgi/input.pl?UserId=pr2yjwXlURmS&sessionId=rLd6tI7IrE5T&input_page_show_search=on
Images and Videos
Cover image: https://www.mdc-berlin.de/16074687/en/news/archive/2008/20080910-erwin_schr_dinger_prize_2008_goes_to_resea/P__PRESS2008_WANKER_SCHROEDINGER_PRIZE_PROTEIN_NETWORK.jpg
Interaction network images: All come from a STRING session.
This website was created for Genetics 564 by Zachary Beethem, an undergraduate genetics major at UW-Madison.
He can be reached via email: [email protected]
Date of last website update: April 2017